Goal¶
To visualize how each metric works and compare them.
Introduction¶
Distance metrics measure how far apart two data points are. They’re fundamental to many ML algorithms like KNN, K-Means clustering, and similarity searches.
we’ll explore:¶
Euclidean Distance - straight-line distance
Manhattan Distance - grid-based distance
Minkowski Distance - generalized distance
Cosine Distance - angle-based similarity
Setup: Import Libraries¶
Source
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.spatial.distance import euclidean, cityblock, cosine, minkowski
from sklearn.metrics.pairwise import euclidean_distances, cosine_distances
# Set style for better visualizations
sns.set_style('whitegrid')
plt.rcParams['figure.figsize'] = (12, 8)
print("✓ All libraries imported successfully!")✓ All libraries imported successfully!
1. Euclidean Distance¶
Formula¶
The straight-line distance between two points. Like measuring distance on a map with a ruler.
Characteristics¶
Most common distance metric
Assumes you can move diagonally
Sensitive to all dimensions equally
Good for most ML tasks
Source
# Example: Calculate Euclidean distance
p = np.array([0, 0])
q = np.array([3, 4])
# Method 1: Manual calculation
dist_manual = np.sqrt((p[0] - q[0])**2 + (p[1] - q[1])**2)
# Method 2: Using scipy
dist_scipy = euclidean(p, q)
print(f"Point P: {p}")
print(f"Point Q: {q}")
print(f"\nEuclidean Distance (Manual): {dist_manual}")
print(f"Euclidean Distance (SciPy): {dist_scipy}")
print(f"\nCalculation: √[(3-0)² + (4-0)²] = √[9 + 16] = √25 = 5")Point P: [0 0]
Point Q: [3 4]
Euclidean Distance (Manual): 5.0
Euclidean Distance (SciPy): 5.0
Calculation: √[(3-0)² + (4-0)²] = √[9 + 16] = √25 = 5
Source
# Visualize Euclidean Distance
fig, ax = plt.subplots(1, 1, figsize=(8, 8))
# Points
ax.plot([0], [0], 'ro', markersize=10, label='Point P (0,0)')
ax.plot([3], [4], 'bs', markersize=10, label='Point Q (3,4)')
# Draw straight line (Euclidean distance)
ax.plot([0, 3], [0, 4], 'r-', linewidth=2, label=f'Euclidean Distance = 5')
# Draw right triangle to show calculation
ax.plot([0, 3], [0, 0], 'gray', linestyle='--', alpha=0.7)
ax.plot([3, 3], [0, 4], 'gray', linestyle='--', alpha=0.7)
# Labels
ax.text(1.5, -0.5, 'Δx = 3', fontsize=12, ha='center')
ax.text(3.5, 2, 'Δy = 4', fontsize=12)
ax.text(1.2, 2.2, 'd = 5', fontsize=14, color='red', fontweight='bold')
ax.set_xlim(-1, 5)
ax.set_ylim(-2, 5)
ax.set_aspect('equal')
ax.grid(True, alpha=0.3)
ax.legend(fontsize=11)
ax.set_xlabel('X-axis', fontsize=12)
ax.set_ylabel('Y-axis', fontsize=12)
ax.set_title('Euclidean Distance: Straight-Line Distance', fontsize=14, fontweight='bold')
plt.tight_layout()
plt.show()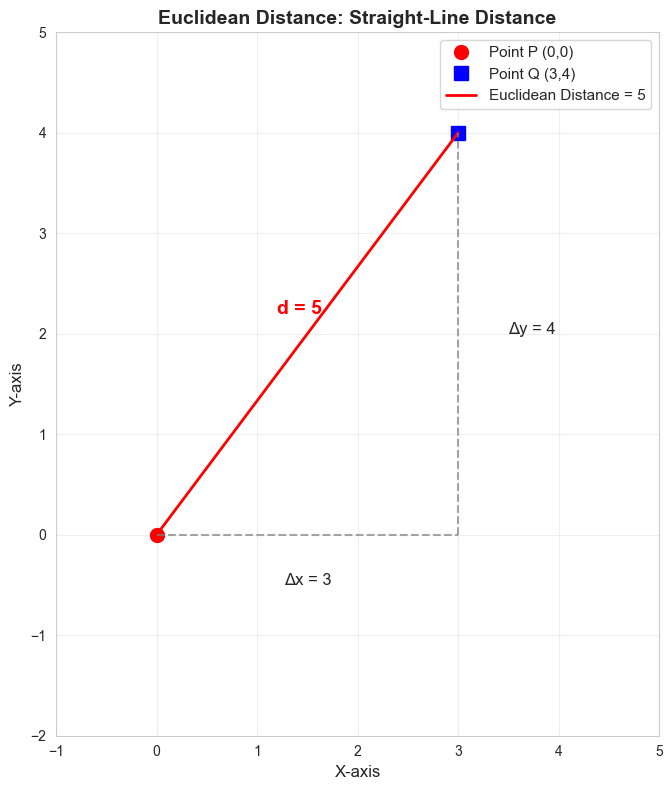
2. Manhattan Distance (L1 Distance)¶
Formula¶
Distance by traveling only horizontally or vertically. Like navigating city blocks.
Characteristics¶
Also called “taxicab distance” or “L1 norm”
You can only move up/down/left/right (no diagonals)
Often faster to compute than Euclidean
More robust to outliers
Source
# Example: Calculate Manhattan distance
p = np.array([0, 0])
q = np.array([3, 4])
# Method 1: Manual calculation
dist_manual = np.abs(p[0] - q[0]) + np.abs(p[1] - q[1])
# Method 2: Using scipy
dist_scipy = cityblock(p, q)
print(f"Point P: {p}")
print(f"Point Q: {q}")
print(f"\nManhattan Distance (Manual): {dist_manual}")
print(f"Manhattan Distance (SciPy): {dist_scipy}")
print(f"\nCalculation: |3-0| + |4-0| = 3 + 4 = 7")
print(f"\nCompare:")
print(f" Euclidean: 5 (straight line)")
print(f" Manhattan: 7 (grid path)")Point P: [0 0]
Point Q: [3 4]
Manhattan Distance (Manual): 7
Manhattan Distance (SciPy): 7
Calculation: |3-0| + |4-0| = 3 + 4 = 7
Compare:
Euclidean: 5 (straight line)
Manhattan: 7 (grid path)
Source
# Visualize Manhattan Distance
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(14, 6))
# Left: One Manhattan path
ax1.plot([0], [0], 'ro', markersize=10, label='Point P (0,0)')
ax1.plot([3], [4], 'bs', markersize=10, label='Point Q (3,4)')
ax1.plot([0, 3], [0, 0], 'g-', linewidth=3, alpha=0.7, label='Move right (3)')
ax1.plot([3, 3], [0, 4], 'b-', linewidth=3, alpha=0.7, label='Move up (4)')
ax1.text(1.5, -0.5, 'Δx = 3', fontsize=12, ha='center')
ax1.text(3.5, 2, 'Δy = 4', fontsize=12)
ax1.set_xlim(-1, 5)
ax1.set_ylim(-2, 5)
ax1.grid(True, alpha=0.3)
ax1.legend(fontsize=10)
ax1.set_title('Manhattan Distance = 7\n(One possible path)', fontsize=12, fontweight='bold')
ax1.set_aspect('equal')
# Right: Multiple Manhattan paths
ax2.plot([0], [0], 'ro', markersize=10, label='Point P (0,0)')
ax2.plot([3], [4], 'bs', markersize=10, label='Point Q (3,4)')
# Path 1: right then up
ax2.plot([0, 3, 3], [0, 0, 4], 'g--', linewidth=2, alpha=0.6)
# Path 2: up then right
ax2.plot([0, 0, 3], [0, 4, 4], 'b--', linewidth=2, alpha=0.6)
ax2.plot([0, 3], [0, 4], 'r-', linewidth=2, label='Euclidean (5)', alpha=0.8)
ax2.text(1.5, 1.5, 'All paths = 7', fontsize=12, color='green', fontweight='bold')
ax2.set_xlim(-1, 5)
ax2.set_ylim(-2, 5)
ax2.grid(True, alpha=0.3)
ax2.legend(fontsize=10)
ax2.set_title('All Manhattan Paths Have Same Distance', fontsize=12, fontweight='bold')
ax2.set_aspect('equal')
plt.tight_layout()
plt.show()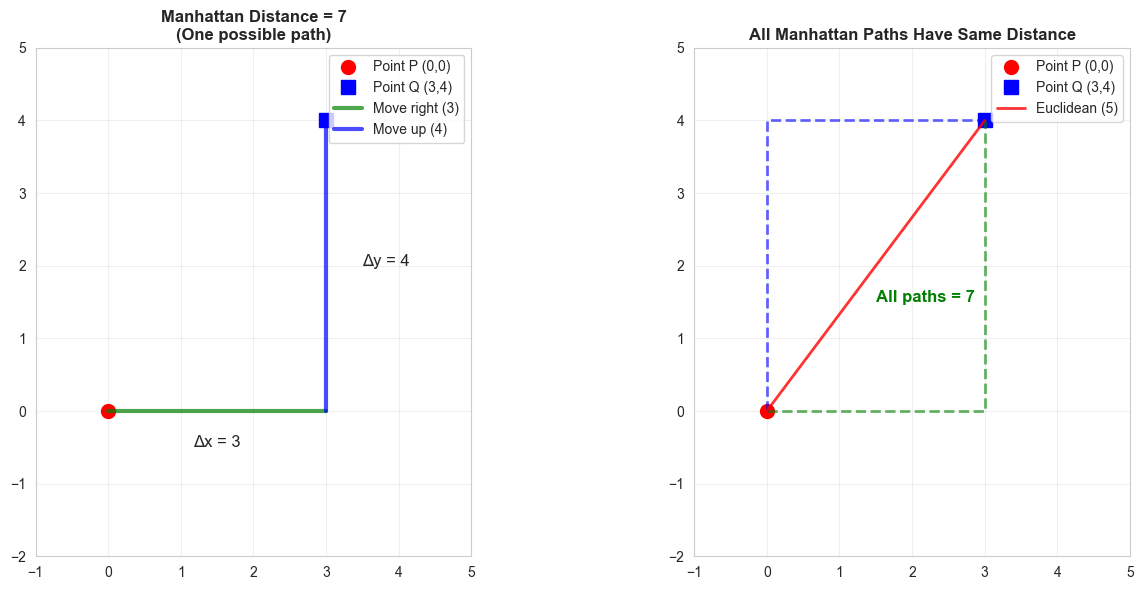
3. Minkowski Distance¶
Formula¶
A generalized distance metric that includes both Euclidean and Manhattan as special cases.
Special Cases¶
p = 1: Manhattan distance
p = 2: Euclidean distance
p = ∞: Chebyshev distance (maximum difference)
Characteristics¶
Flexible - can adjust the parameter p
p=1 emphasizes absolute differences
p=2 is most commonly used
p>2 emphasizes larger differences
Source
# Example: Calculate Minkowski distance for different p values
p_point = np.array([0, 0])
q_point = np.array([3, 4])
print(f"Point P: {p_point}")
print(f"Point Q: {q_point}")
print(f"\nMinkowski Distance for different p values:")
print("="*50)
for p in [1, 1.5, 2, 3, np.inf]:
if p == np.inf:
dist = np.max(np.abs(p_point - q_point))
print(f"p = ∞ (Chebyshev): {dist}")
else:
dist = minkowski(p_point, q_point, p)
print(f"p = {p:<3} (L{p} norm): {dist:.4f}")Point P: [0 0]
Point Q: [3 4]
Minkowski Distance for different p values:
==================================================
p = 1 (L1 norm): 7.0000
p = 1.5 (L1.5 norm): 5.5843
p = 2 (L2 norm): 5.0000
p = 3 (L3 norm): 4.4979
p = ∞ (Chebyshev): 4
Source
# Visualize how p affects Minkowski distance
p_values = np.linspace(1, 5, 50)
distances = [minkowski(p_point, q_point, p) for p in p_values]
plt.figure(figsize=(10, 6))
plt.plot(p_values, distances, 'b-', linewidth=2.5)
plt.axhline(y=5, color='r', linestyle='--', linewidth=2, alpha=0.7, label='Euclidean (p=2) = 5')
plt.axhline(y=7, color='g', linestyle='--', linewidth=2, alpha=0.7, label='Manhattan (p=1) = 7')
plt.scatter([1, 2], [7, 5], s=100, c=['green', 'red'], zorder=5)
plt.xlabel('p (parameter)', fontsize=12)
plt.ylabel('Distance', fontsize=12)
plt.title('Minkowski Distance Changes with p Parameter', fontsize=14, fontweight='bold')
plt.grid(True, alpha=0.3)
plt.legend(fontsize=11)
plt.tight_layout()
plt.show()
print("Key insight: As p increases, Minkowski distance decreases toward Chebyshev distance")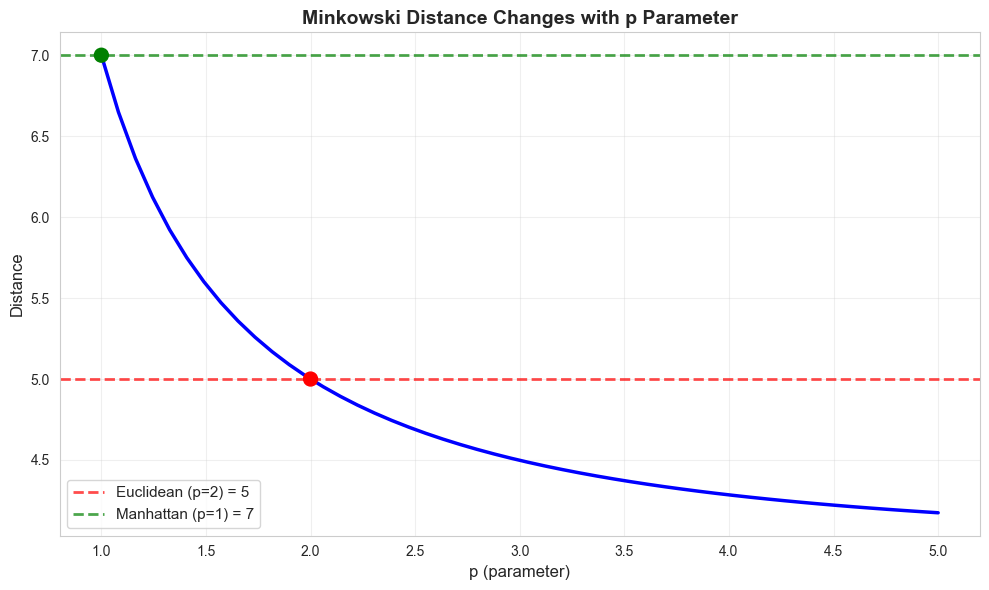
Key insight: As p increases, Minkowski distance decreases toward Chebyshev distance
Source
# Example: Calculate Cosine distance
p = np.array([1, 0])
q = np.array([1, 1])
# Method 1: Manual calculation
dot_product = np.dot(p, q)
magnitude_p = np.linalg.norm(p)
magnitude_q = np.linalg.norm(q)
cosine_sim = dot_product / (magnitude_p * magnitude_q)
cosine_dist = 1 - cosine_sim
# Method 2: Using scipy
cosine_dist_scipy = cosine(p, q)
print(f"Point P: {p}")
print(f"Point Q: {q}")
print(f"\nDot product: {dot_product}")
print(f"Magnitude of P: {magnitude_p:.4f}")
print(f"Magnitude of Q: {magnitude_q:.4f}")
print(f"\nCosine Similarity: {cosine_sim:.4f}")
print(f"Cosine Distance (Manual): {cosine_dist:.4f}")
print(f"Cosine Distance (SciPy): {cosine_dist_scipy:.4f}")
print(f"\nInterpretation: Vectors point in similar directions (angle ≈ 45°)")Point P: [1 0]
Point Q: [1 1]
Dot product: 1
Magnitude of P: 1.0000
Magnitude of Q: 1.4142
Cosine Similarity: 0.7071
Cosine Distance (Manual): 0.2929
Cosine Distance (SciPy): 0.2929
Interpretation: Vectors point in similar directions (angle ≈ 45°)
Source
# Visualize Cosine Distance
fig, axes = plt.subplots(1, 3, figsize=(15, 5))
# Case 1: Similar direction (low cosine distance)
ax = axes[0]
p1, q1 = np.array([1, 0.5]), np.array([2, 1])
ax.arrow(0, 0, p1[0], p1[1], head_width=0.1, head_length=0.1, fc='red', ec='red', linewidth=2, label='Vector P')
ax.arrow(0, 0, q1[0], q1[1], head_width=0.1, head_length=0.1, fc='blue', ec='blue', linewidth=2, label='Vector Q')
cos_sim1 = cosine(p1, q1)
ax.set_xlim(-0.5, 3)
ax.set_ylim(-0.5, 2)
ax.set_aspect('equal')
ax.grid(True, alpha=0.3)
ax.legend(fontsize=11)
ax.set_title(f'Similar Direction\nCosine Distance = {1-cos_sim1:.4f}', fontsize=12, fontweight='bold')
# Case 2: Perpendicular (cosine distance = 1)
ax = axes[1]
p2, q2 = np.array([1, 0]), np.array([0, 1])
ax.arrow(0, 0, p2[0], p2[1], head_width=0.1, head_length=0.1, fc='red', ec='red', linewidth=2, label='Vector P')
ax.arrow(0, 0, q2[0], q2[1], head_width=0.1, head_length=0.1, fc='blue', ec='blue', linewidth=2, label='Vector Q')
cos_sim2 = cosine(p2, q2)
ax.set_xlim(-0.5, 1.5)
ax.set_ylim(-0.5, 1.5)
ax.set_aspect('equal')
ax.grid(True, alpha=0.3)
ax.legend(fontsize=11)
ax.set_title(f'Perpendicular (90°)\nCosine Distance = {1-cos_sim2:.4f}', fontsize=12, fontweight='bold')
# Case 3: Opposite direction (cosine distance = 2)
ax = axes[2]
p3, q3 = np.array([1, 0]), np.array([-1, 0])
ax.arrow(0, 0, p3[0], p3[1], head_width=0.1, head_length=0.1, fc='red', ec='red', linewidth=2, label='Vector P')
ax.arrow(0, 0, q3[0], q3[1], head_width=0.1, head_length=0.1, fc='blue', ec='blue', linewidth=2, label='Vector Q')
cos_sim3 = cosine(p3, q3)
ax.set_xlim(-1.5, 1.5)
ax.set_ylim(-0.5, 0.5)
ax.set_aspect('equal')
ax.grid(True, alpha=0.3)
ax.legend(fontsize=11)
ax.set_title(f'Opposite Direction (180°)\nCosine Distance = {1-cos_sim3:.4f}', fontsize=12, fontweight='bold')
plt.tight_layout()
plt.show()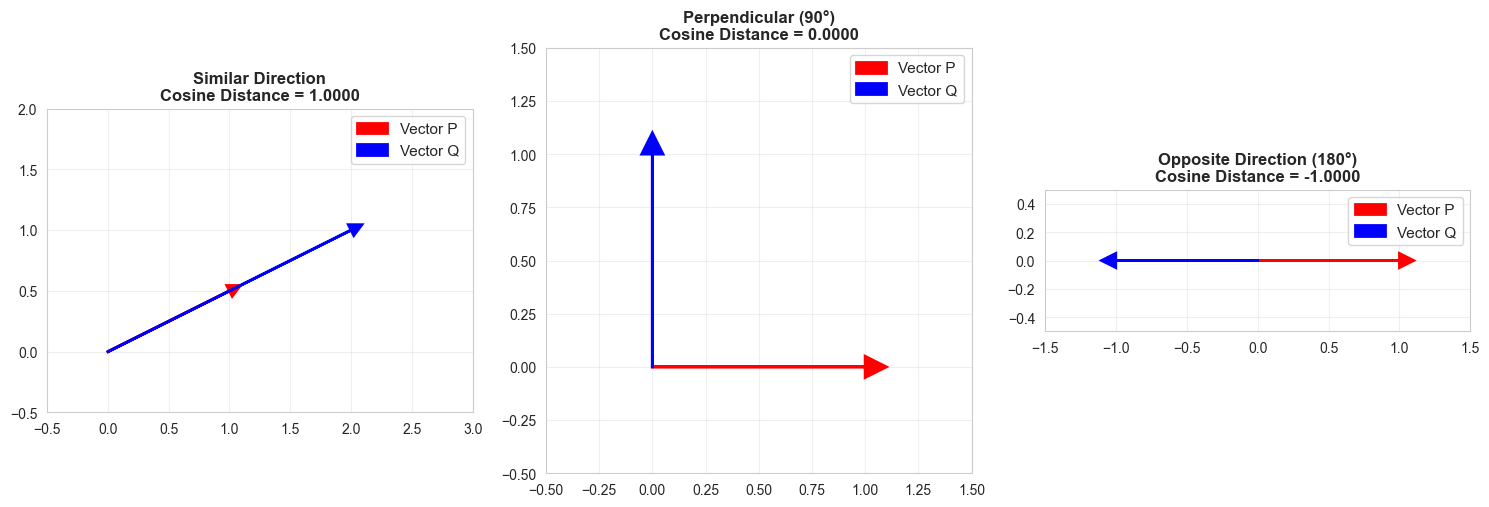
5. Comparison: All Distance Metrics¶
Source
# Create a comparison table
p = np.array([0, 0])
q = np.array([3, 4])
metrics = {
'Euclidean': euclidean(p, q),
'Manhattan': cityblock(p, q),
'Minkowski (p=3)': minkowski(p, q, 3),
'Chebyshev': np.max(np.abs(p - q)),
'Cosine': cosine(p, q)
}
print("\nDistance Metric Comparison")
print(f"Point P: {p}, Point Q: {q}")
print("="*50)
for metric, distance in metrics.items():
print(f"{metric:<20} {distance:.4f}")
Distance Metric Comparison
Point P: [0 0], Point Q: [3 4]
==================================================
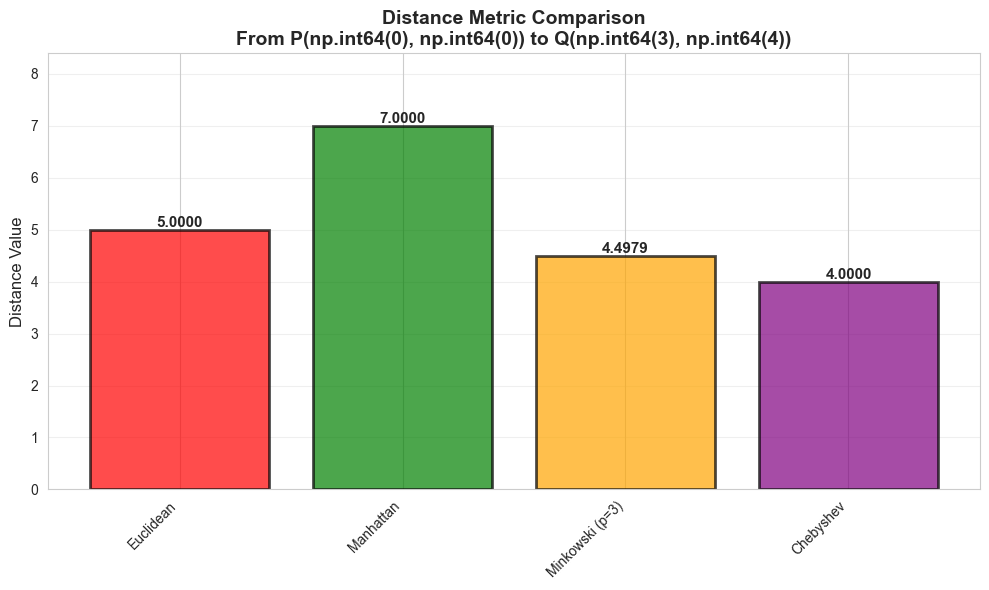
Euclidean 5.0000
Manhattan 7.0000
Minkowski (p=3) 4.4979
Chebyshev 4.0000
Cosine nan
/Users/deepakkandalam/workspace/datawizard.github.io/.venv/lib/python3.9/site-packages/scipy/spatial/distance.py:647: RuntimeWarning: invalid value encountered in divide
dist = 1.0 - uv / math.sqrt(uu * vv)
Source
# Visual comparison
metric_names = list(metrics.keys())
metric_values = list(metrics.values())
plt.figure(figsize=(10, 6))
bars = plt.bar(metric_names, metric_values, color=['red', 'green', 'orange', 'purple', 'blue'], alpha=0.7, edgecolor='black', linewidth=2)
# Add value labels on bars
for bar, value in zip(bars, metric_values):
height = bar.get_height()
plt.text(bar.get_x() + bar.get_width()/2., height,
f'{value:.4f}', ha='center', va='bottom', fontsize=11, fontweight='bold')
plt.ylabel('Distance Value', fontsize=12)
plt.title(f'Distance Metric Comparison\nFrom P{tuple(p)} to Q{tuple(q)}', fontsize=14, fontweight='bold')
plt.ylim(0, max(metric_values) * 1.2)
plt.xticks(rotation=45, ha='right')
plt.grid(True, alpha=0.3, axis='y')
plt.tight_layout()
plt.show()posx and posy should be finite values
posx and posy should be finite values
posx and posy should be finite values
6. When to Use Each Metric¶
| Metric | Best For | Pros | Cons |
|---|---|---|---|
| Euclidean | Most ML tasks, KNN | Most intuitive, works well generally | Sensitive to outliers |
| Manhattan | High-dimensional data, robust cases | Faster, robust to outliers | Less intuitive |
| Minkowski | Flexible needs, parameter tuning | Can adjust for different problems | More complex |
| Cosine | Text/document similarity, sparse data | Direction-based, ignores scale | Not true distance |
| Chebyshev | Game algorithms, grid-based problems | Fast to compute | Less common in ML |
Decision Guide:¶
Default choice: Euclidean distance
Text/NLP: Cosine distance
High dimensions: Manhattan or Cosine
Need robustness: Manhattan
Grid-based problems: Chebyshev
Source
# Real-world example: Distance between movie ratings
# User 1 rated: Action=5, Comedy=3, Drama=4
# User 2 rated: Action=5, Comedy=4, Drama=3
user1 = np.array([5, 3, 4])
user2 = np.array([5, 4, 3])
print("\n" + "="*60)
print("Real-World Example: Movie Rating Similarity")
print("="*60)
print(f"\nUser 1 ratings [Action, Comedy, Drama]: {user1}")
print(f"User 2 ratings [Action, Comedy, Drama]: {user2}")
euc_dist = euclidean(user1, user2)
cos_sim = cosine(user1, user2)
print(f"\nEuclidean Distance: {euc_dist:.4f}")
print(f" → Ratings are {euc_dist:.2f} units apart")
print(f" → Users have different rating patterns")
print(f"\nCosine Distance: {cos_sim:.4f}")
print(f" → Their rating directions are very similar (cosine sim = {1-cos_sim:.4f})")
print(f" → Both users like similar types of movies (just different ratings)")
print(f"\nConclusion:")
print(f" → Use Euclidean if you care about magnitude of differences")
print(f" → Use Cosine if you only care about preference direction")
============================================================
Real-World Example: Movie Rating Similarity
============================================================
User 1 ratings [Action, Comedy, Drama]: [5 3 4]
User 2 ratings [Action, Comedy, Drama]: [5 4 3]
Euclidean Distance: 1.4142
→ Ratings are 1.41 units apart
→ Users have different rating patterns
Cosine Distance: 0.0200
→ Their rating directions are very similar (cosine sim = 0.9800)
→ Both users like similar types of movies (just different ratings)
Conclusion:
→ Use Euclidean if you care about magnitude of differences
→ Use Cosine if you only care about preference direction
Summary¶
✅ Euclidean: Default choice, straight-line distance
✅ Manhattan: Grid-based, robust alternative
✅ Minkowski: Flexible generalization (p=1 → Manhattan, p=2 → Euclidean)
✅ Cosine: Angle-based, great for text/vectors
Key Takeaways:¶
Different metrics measure distance differently
Choice of metric affects KNN, clustering, and other algorithms
Euclidean is the safest default
Cosine is best for high-dimensional and text data
Always test multiple metrics for your specific problem